Overview
Airborne light detection and ranging (lidar) data have great potential to map vegetation structure at a fine resolution, including estimating carbon stocks, canopy closure, and tree height. While airborne-lidar-derived vegetation structural measurements could play an important role in ecological restoration monitoring, these data are limited to only a tiny fraction of the Earth’s surface. Data fusion to combine high-resolution lidar data with medium resolution satellite data (such as Landsat imagery) represents one way to extend the spatial and temporal reach of airborne data.
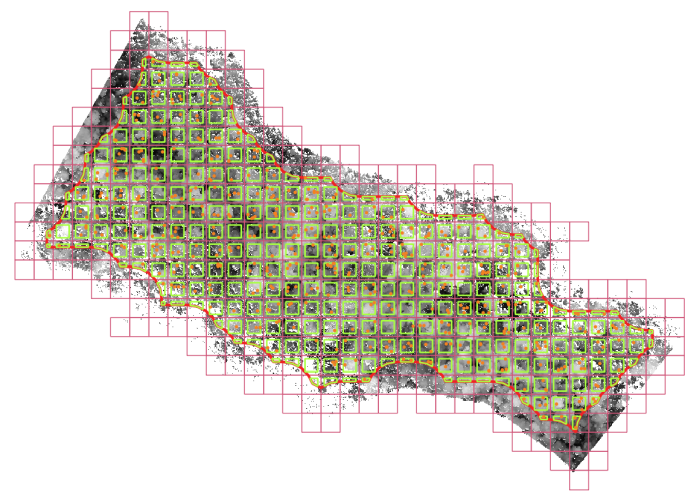
This data set includes 1,105 aerial images captured over Mexico as part of the NASA G-LiHT program, along with lidar-derived canopy height values and Landsat-derived vegetation indices for 499,925 sample points within those images.
The associated open-source repo provides tools required to replicate this sampling methodology.
Data format
G-LiHT images are in GeoTIFF format.
Annotations are provided in a .csv file with 499925 rows, each representing one sample, and 12 columns:
- EVI: Landsat-derived enhanced vegetation index
- NVDI: Landsat-derived normalized difference vegetation index
- NDWI: Landsat-derived normalized difference water index
- NBR: Landsat-derived normalized burn ratio
- CanopyHeig: lidar-derived canopy height in meters
- Ecoregion: Ecoregion label, e.g. “Tropical and Subtropical Coniferous Forests”
- ID_G_LiTH_: G-LiHT ID image; corresponds to the filenames in the “Images” folder
- ID_Landsat: Unique ID for each Landsat pixel
- X: longitude
- Y: latitude
- Vegetation: vegetation type extracted from the Mexican National Institute of Statistics and Geography (INEGI), e.g. “AGRICULTURA DE TEMPORAL ANUAL”
- field_1: field identifier (a unique row identifier)
Citation
If you use these data in a publication or report, please use the following citation:
Requena-Mullor JM, Caughlin TT. Lidar-derived canopy height data and Landsat-derived vegetation indices for the G-LiHT NASA acquisitions in Mexico. Dataset.
Contact information
For questions about this data set, contact Juan Miguel Requena Mullor (juanmimullor@gmail.com).
License
G-LiHT images are released according to NASA’s Data and Information Policy.
The .csv file containing canopy height and vegetation information is released under the Community Data License Agreement (permissive variant).
Download links
Metadata (GCP link, 21MB)
Metadata (Azure link, 21MB)
Images (62.9GB) are available in the following cloud storage folders. See LILA’s direct image access guide for download instructions.
Google Cloud Storage folder
gs://public-datasets-lila/boise-state-vegetation/tifImages
Azure Blob Storage folder
https://lilablobssc.blob.core.windows.net/boise-state-vegetation/tifImages
Having trouble downloading? Check out our FAQ.
Posted by Dan Morris.