Overview
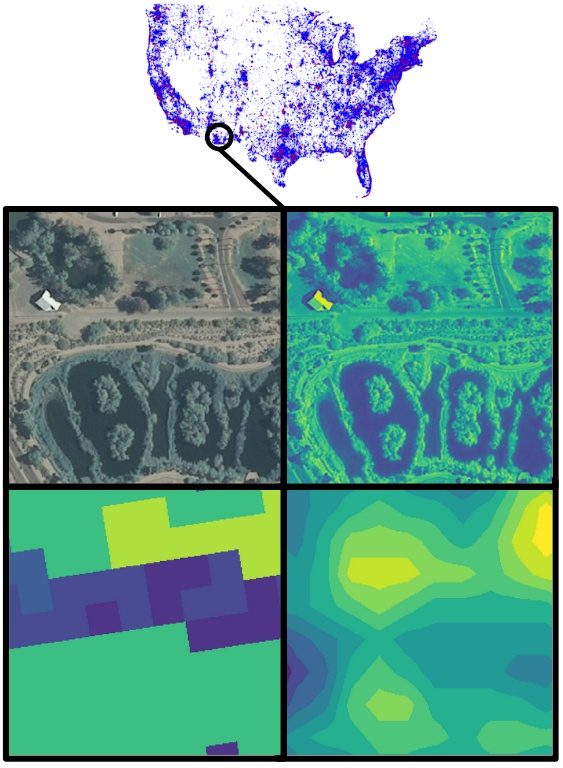
Understanding the geographic distribution of species is a key concern in conservation. By pairing species occurrences with environmental features, researchers can model the relationship between an environment and the species which may be found there. To facilitate research in this area, we present the GeoLifeCLEF 2020 (GLC2020) dataset, which consists of 1.9 million geo-located species observations paired with high-resolution remote sensing imagery, land cover data, and altitude, in addition to traditional low-resolution climate and soil variables. The observations in this dataset cover 31,435 plant and animal species from the United States and France. The dataset was originally prepared for the GeoLifeCLEF 2020 competition. Full details can be found in the dataset paper (cited below).
Citation
If you use this data set, please cite the associated manuscript:
Cole E, Deneu B, Lorieul T, Servajean M, Botella C, Morris D, Jojic N, Bonnet P, Joly A. The GeoLifeCLEF 2020 Dataset. arXiv preprint arXiv:2004.04192. 2020 Apr 8. (bibtex)
Data format
The dataset consists of three components: high-resolution patches, annotations for those patches, and low-resolution covariate rasters.
Each high-resolution patch is stored as a pair of .npy files: XXX.npy containing RGB-IR and land cover (256x256x5 uint8 array) and XXX_alti.npy containing altitude (256x256x1 uint16 array).
Annotations for the high-resolution patches are provided as JSON files that adhere to a modified version of the COCO dataset annotation format. The format is as follows:
{
"images": [image],
"categories": [category],
"annotations": [annotation]
}
image
{
"id": int,
"width": int,
"height": int,
"file_name": str,
"file_name_alti": str,
"lon": float,
"lat": float,
"country": str
}
category
{
"id": int,
"gbif_id": int,
"gbif_name": str
}
annotation
{
"id": int,
"image_id": int,
"category_id" int
}
The low-resolution covariate rasters are provided as pairs of .tif files (one for the US and one for France) for each variable. On the competition GitHub page we provide code to extract values for 27 environmental characteristics at any location in the US or France.
Download links
High-resolution-patches (GCP links)
High-resolution patches are available at:
https://storage.googleapis.com/public-datasets-lila/geolifeclef-2020/patches_[region]_[index].tar.gz (11GB each)
…where region is “us” or “fr” for the US and France, respectively, and “index” is 01, 02, … 20. For example:
https://storage.googleapis.com/public-datasets-lila/geolifeclef-2020/patches_fr_01.tar.gz
High-resolution-patches (AWS links)
High-resolution patches are available at:
http://us-west-2.opendata.source.coop.s3.amazonaws.com/agentmorris/lila-wildlife/geolifeclef-2020/patches_[region]_[index].tar.gz (11GB each)
…where region is “us” or “fr” for the US and France, respectively, and “index” is 01, 02, … 20. For example:
High-resolution-patches (Azure links)
High-resolution patches are available at:
https://lilawildlife.blob.core.windows.net/lila-wildlife/geolifeclef-2020/patches_[region]_[index].tar.gz (11GB each)
…where region is “us” or “fr” for the US and France, respectively, and “index” is 01, 02, … 20. For example:
https://lilawildlife.blob.core.windows.net/lila-wildlife/geolifeclef-2020/patches_fr_01.tar.gz
High-resolution-patches (folder structure)
For each country, each of the 20 .tar.gz files contains five directories from the set {00/, …, 99/}. For example, patches_fr_01.tar.gz contains 00/, …, 04/ and patches_fr_02.tar.gz contains 05/, …, 09/, and so on. Then each of those directories contains subdirectories 00/, …, 99/. These indices do not have a semantic interpretation, they are just to help the operating system deal with the number of files. Each pair of patch files is named XXXXABCD.npy and XXXXABCD_alti.npy, and lives at CD/AB/ where CD and AB are each among 00/, …, 99/.
The contents of each file should be extracted to the same directory, such that, for example, the 00/ folder contains the contents of both patches_us_01/ and patches_fr_01/.
Annotations
- Training (450MB) (GCP link) (Azure link) (AWS link)
- Validation (11MB) (GCP link) (Azure link) (AWS link)
- Testing (13MB) (GCP link) (Azure link) (AWS link)
Low-resolution covariate rasters
Covariate rasters (2.4GB) (GCP link) (Azure link) (AWS link)
Having trouble downloading? Check out our FAQ.
Contact information
For questions about this dataset, contact geolifeclef@inria.fr.
License information
RGB-IR imagery
US
Source: NAIP/USGS
License: public domain
FR
Source: IGN
License: CC BY 4.0 (By Permission of IGN)
Land cover
US
Source: NLCD/USGS
License: public domain
FR
Source: CESBIO
License: ODC-BY 1.0
Species occurrences
US
Source: iNaturalist
License: CC BY-NC 4.0
FR
Source #1: Pl@ntNet and iNaturalist
License #1: CC BY 4.0
Source #2: iNaturalist
License #2: CC BY-NC 4.0
Altitude
Source: SRTM/NASA
License: public domain
Soil rasters
Source: SoilGrids
License: CC BY 4.0
Bioclimatic rasters
Source: WorldClim
License: CC BY-SA 4.0
Posted by Dan Morris.